The Hyve has developed a forest plot for the variant page in Open Targets Genetics. A forest plot, which has been included in the recently released version V8 - 22.10, allows researchers to compare traits among different groups of phenotypes, helping them to find new insights or outliers relevant to their research.
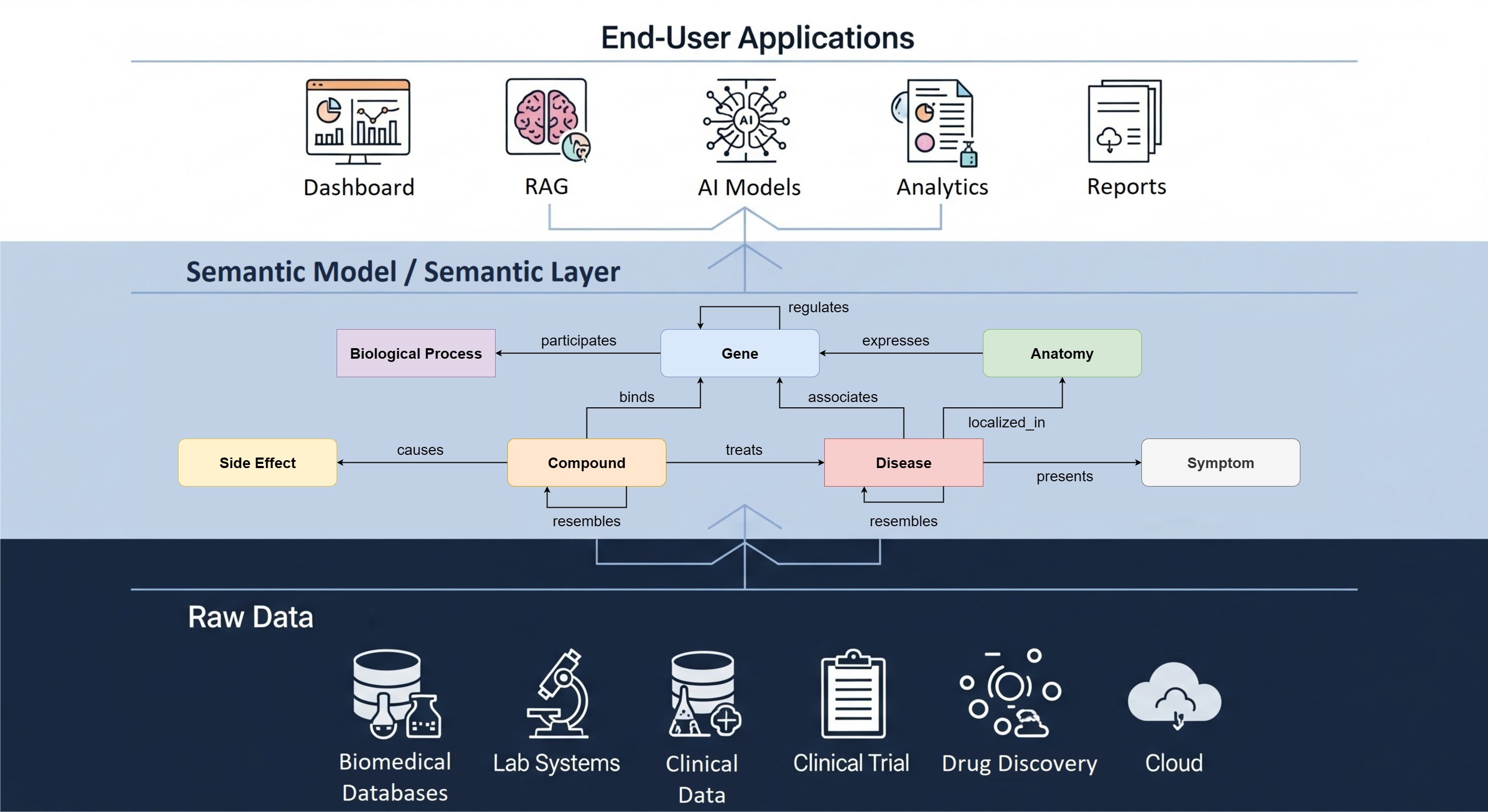
Open Targets Genetics is a powerful solution for both GWAS data processing, integration, and visualization. It integrates GWAS studies from sources such as UK Biobank and FinnGen. Multiple methods are used to analyze these GWAS studies. Fine-mapping across thousands of traits is done to extract significant variants, colocalization analysis is performed to compare against other studies, and variants are linked to target genes using a Locus2Gene assessment. All the data from GWAS studies and analysis pipelines are visualized in the Open Targets Genetics web application. For more information about Open Targets Genetics can be found here.
The Variant Page
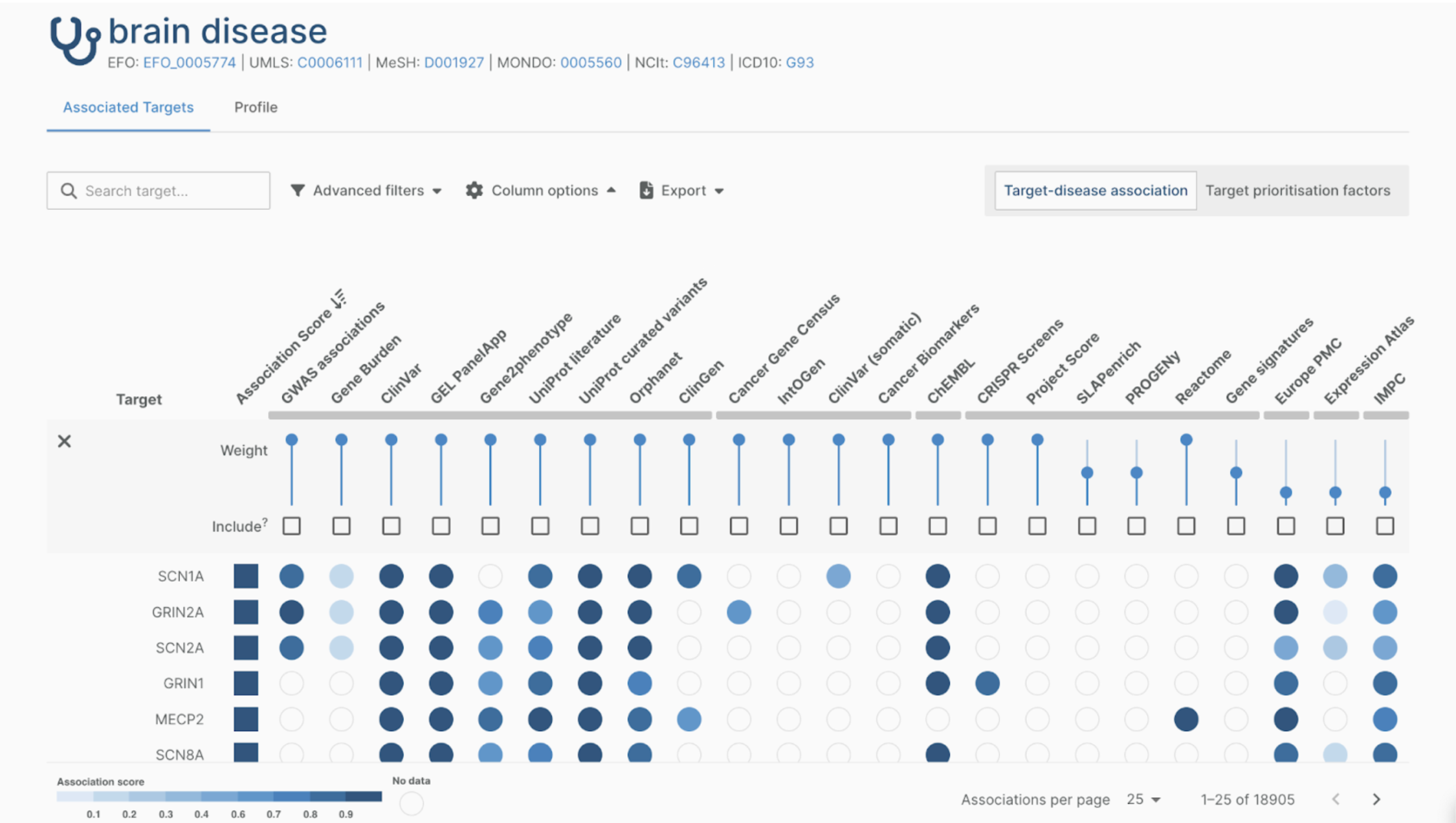
As described before, the data from GWAS studies and analysis pipelines is visualized in the Open Targets Genetics web application. This web interface consists of multiple pages, like the Study, Gene, and Variant pages. Our contribution to the newly released Open Targets Genetics version is the addition of a forest plot located on the Variant page. So let’s take a look at the Variant page.
The Variant page consists of five sections. The assigned genes, PheWAS, Forest Plot, lead variants, and tag variants section. As the name suggests, the assigned genes section shows which genes are functionally implicated by the variant. The lead variants and tag variants sections show which lead and tag variants are linked to the variant. The PheWAS section shows which traits are associated with the variant through a PheWAS plot. Finally, the Forest Plot section contains the newly implemented forest plot and a table containing the associated traits.
The PheWAS plot shows − through p-values − how likely it is that a particular variant is associated with specific traits. These traits are color-coded by trait category to make it easier for researchers to compare categories with each other. However, this PheWAS plot does not show the effect sizes and lacks the functionality to focus on trait categories of interest. One of our clients requested us to implement a forest plot that can be used next to the PheWAS plot, and that does have this functionality. By implementing this forest plot, their scientists are able to better analyze the data.
Introducing the forest plot
The forest plot has the benefit that it provides both tabular and graphical information. In the table on the left-hand side, trait names and p-values are shown. The graphical representation on the right-hand side shows effect sizes with their corresponding 95% confidence intervals. The traits are vertically aligned, grouped, and color-coded by trait category and ordered by effect size.

How can I use the forest plot?
One use case for the forest plot in Open Targets Genetics is finding traits within one or more trait category groups associated with a variant of interest. If there is a significant association between the variant and a trait, this variant could be a target for drug development toward the trait of interest.
Let's take a look at an example of this use case.
First, we select a variant of interest that is associated with the waist-to-hip ratio adjusted for BMI by following the steps described below:
Search for “Waist-to-hip ratio adjusted for BMI”
Select the fifth study: https://genetics.opentargets.org/study/GCST008733 (author: Lotta LA, study_id: GCST008733)
From the Independently-associated loci table, select the variant with the lowest p-value: https://genetics.opentargets.org/variant/6_43790159_C_A (6_43790159_C_A)
Now let’s identify what other traits this variant is associated with:
- From the PheWAS plot, we will identify trait categories of interest. The ‘cardiovascular disease,’ ‘phenotype,’ and ‘pancreas disease’ trait categories seem to have a high number of significant traits.

- Let’s select these trait categories in the dropdown of the forest plot, which will give you the forest plot shown below.

As you can see, filtering the number of trait categories makes it much easier to focus on specific groups of traits. The effect size shows the magnitude and direction of the effect, which helps to identify traits of interest.
Open Targets services by The Hyve
Interested in having your own Open Targets installations to analyze your own data in a familiar interface? Or just curious to see what services The Hyve provides around Open Targets? Find out more about our Open Targets services and our custom Open Target Genetics instance.