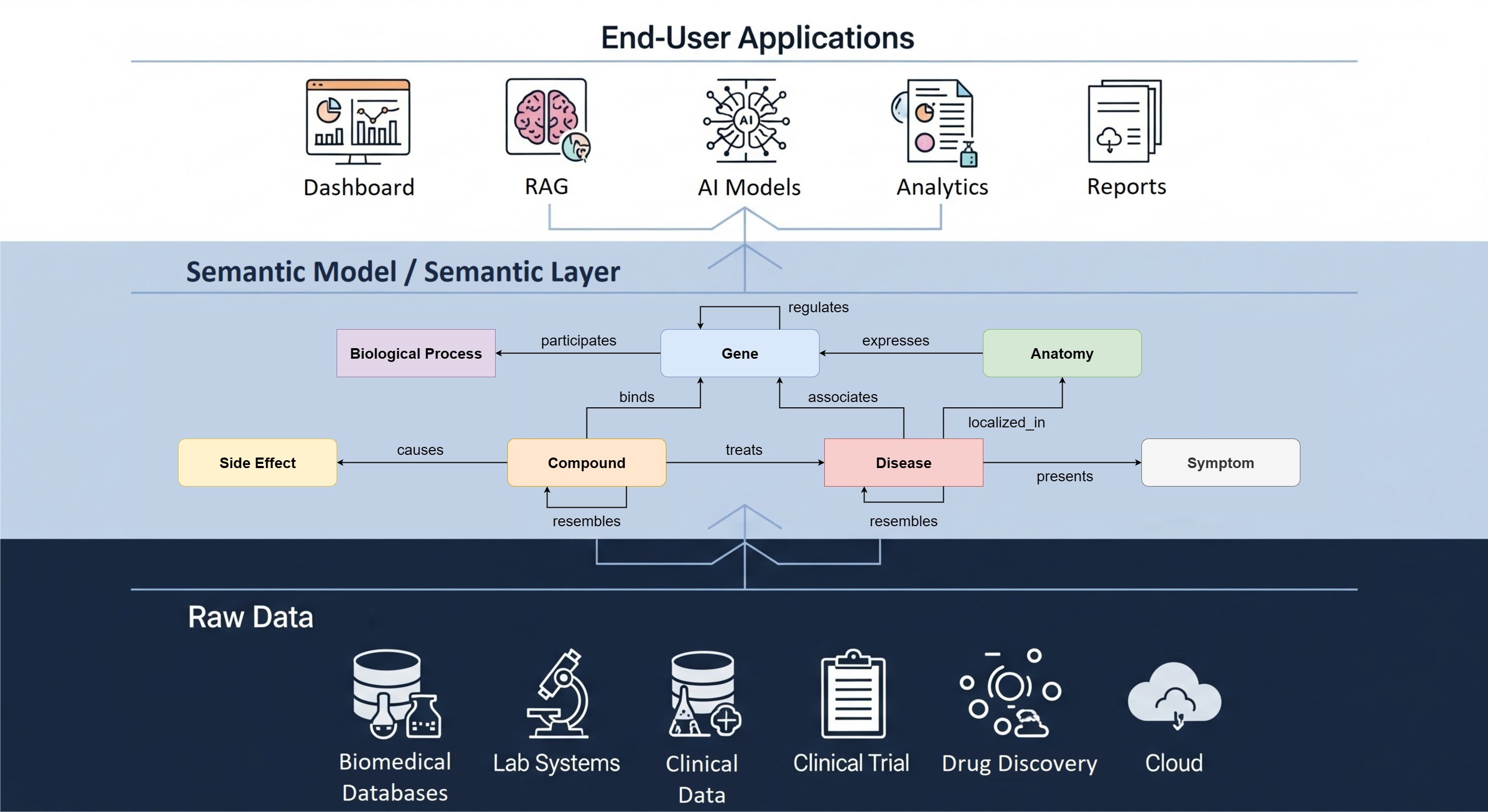
To simplify extraction of a specific data subset from a tranSMART data warehouse for analysis with Python or R, The Hyve developed both a tranSMART Python and a R client.
tranSMART is widely used by pharmaceutical companies, universities, and academic hospitals for storage, sharing and analysis of patient and biomedical research data.
The data warehouse was developed in 2009 by Janssen R&D and subsequently open-sourced in 2012.
Latest developments
In the past two years, The Hyve has developed tranSMART 17.1, that incorporated new functionalities such as the ability to follow patients over time and cross study analysis of different of samples types, for example, the comparison of healthy and tumour tissue. The new tranSMART version also allows for queries to be saved and re-executed. This project was sponsored by Pfizer, Sanofi, Abbvie and Roche.
At the same time, The Hyve developed Glowing Bear. This new tool enables scientists to select specific patient and sample cohorts from the tranSMART 17.1 database that could not be identified before, such as selection of patients based on year of treatment.
tranSMART and Python
Users of the TranSMART platform will know that it has a range of data analysis and visualization options (tools?) built-in. More advanced users prefer to write their own custom analyses using Python or R. To make life easier for them, we developed a Python client and a R client application that speeds up the process of importing selected subsets of data from tranSMART directly into Python or R.
The tranSMART client applications are especially helpful when working with Glowing Bear for the selection of patient subsets and sample cohorts. Previously, you needed to export a subset of data in the right format, download it to your computer and load it into the application of choice. With the client application, this can all be done in one go.